When researchers test whether an antibiotic will work, they usually do so in a controlled laboratory environment. But when an infection happens inside the human body, things aren’t so clean and tidy. New research from the Levin Lab published in mBio, an American Society for Microbiology journal, found that even a slight change in acidity may dramatically shift how bacteria respond to treatment.
The study is centered around Klebsiella pneumoniae, a major cause of deadly infections and one of the world’s most antibiotic-resistant pathogens. “We’re very rapidly approaching a point where antibiotic resistance is going to be a real problem. We have to work on ways of maximizing what we already have, making our arsenal of drugs more effective,” said Sarah Beagle, staff scientist and lead author on the study. “It’s going to be very critical moving forward.”
So, the Levin Lab set out to learn what happens to antibiotic resistance when K. pneumoniae grows in mildly acidic conditions like those found in parts of the human body during an active infection. What they found was that when grown at a pH of 5, the bacterium became up to 64 times more resistant to beta-lactam antibiotics, the most widely prescribed treatment for infections.
These beta-lactams work by shutting down the cell wall-building proteins in the bacterium known as PBPs. Without these, the bacterium can’t properly construct its cell wall or divide, and it eventually dies. However, in addition to these PBPs made at neutral pH, it appears that K. pneumoniae has a reserve team ready to step up when conditions get acidic.
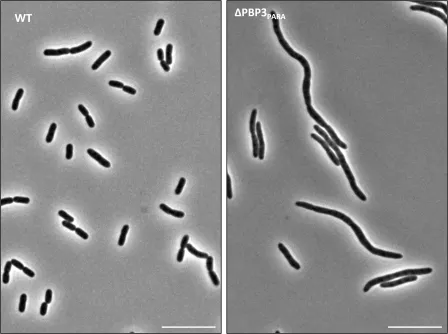
“K. pneumoniae has a backup set of cell wall-building proteins that come online as the cell enters, in this case, an acidic environment,” explained Beagle. PBP2PARA and PBP3PARA are duplicate copies of essential cell wall synthesis genes, and when conditions turn acidic, these alternate versions of cell building and division proteins are expressed. This was surprising, given that past research on pathogen resistance has been done on a model organism, E. coli, which does not have a backup team of proteins. “This sets up a lot of implications for reevaluating and rethinking how we’re assessing antibiotic resistance in pathogens,” said Beagle.
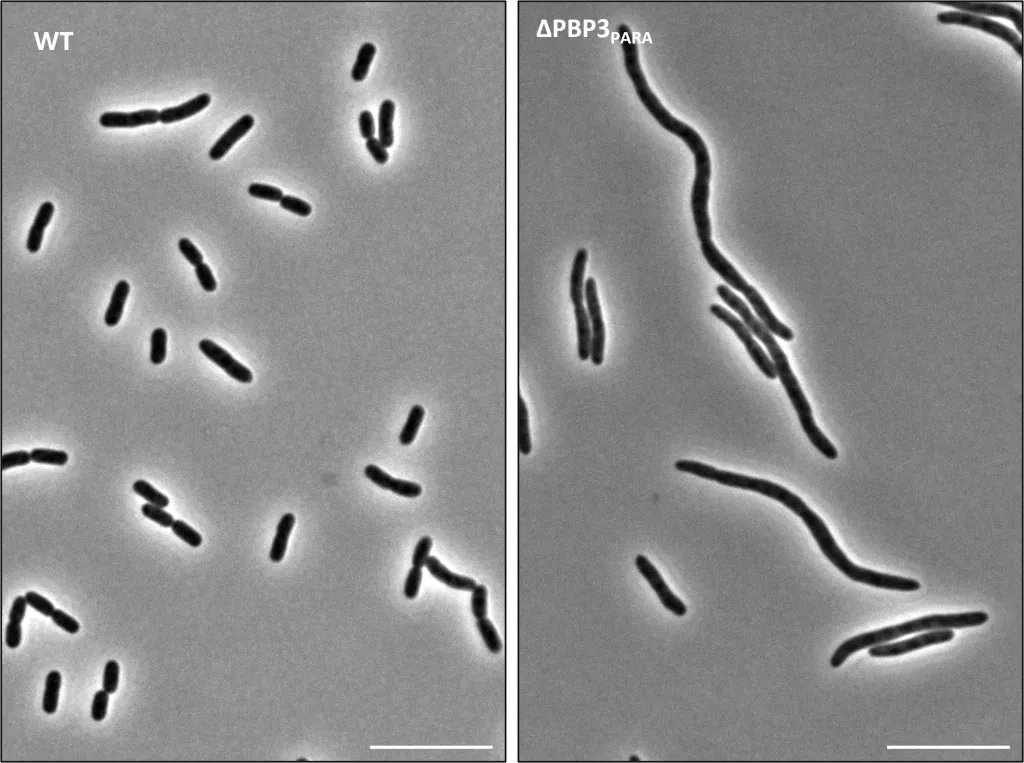
Additionally, they identified another duplicate cell wall synthesis protein, PBP1b, whose activity appeared to be important for stress response during growth at low pH. These results suggest that these duplicate proteins may be important for helping the bacterium survive against antibiotic treatments under acidic conditions.
To confirm this, researchers silenced these proteins. “The approach for studying genetics in bacteria is, if you want to study it, you have to break it,” said Beagle. And so they did, and they found that when these proteins weren’t made, the cell lost much of its antibiotic resistance. “PBP1b and PBP3PARA make the most impact on resistance, so their presence is most critical to the cell at low pH,” confirmed Beagle.
In the face of antibiotic resistance, these findings offer a warning and a potential path forward. “I’m hoping this research leads to a reevaluation of the way we screen for novel drugs,” said Beagle.
In the future, the Levin Lab is looking to widen the kinds of conditions they are screening antibiotic sensitivity in, beyond changes in pH. By designing experiments that better reflect the realities of infection, scientists may be able to extend the usefulness of existing antibiotics and develop combination strategies that are more effective where it matters most: “The ultimate goal is to look for compounds or therapeutics that we can use alongside our current antibiotics to kill more effectively under host-like conditions.”
This research was funded by the National Institutes of Health (5R35GM127331).